I-Nat Med | Indlela ye-multi-omics yokumaka indawo ehlanganisiwe yesimila, amasosha omzimba kanye namagciwane omdlavuza we-colorectal yembula ukusebenzisana kwe-microbiome nesistimu yomzimba yokuzivikela.
Nakuba ama-biomarker omdlavuza wamathumbu amakhulu aye afundwa kabanzi eminyakeni yamuva nje, iziqondiso zamanje zemitholampilo zithembele kuphela ekuhleleni nasekutholeni amaphutha okulungisa ukungalingani kwe-DNA (MMR) noma ukungazinzile kwe-microsatellite (MSI) (ngaphezu kokuhlolwa okujwayelekile kwe-pathology) ukuze kunqunywe izincomo zokwelapha. Abacwaningi baphawule ukuntuleka kokuxhumana phakathi kwezimpendulo zomzimba ezisekelwe ekubonakalisweni kwezakhi zofuzo, amaphrofayili e-microbial, kanye ne-tumor stroma eqenjini lomdlavuza we-colorectal we-Cancer Genome Atlas (TCGA) kanye nokusinda kwesiguli.
Njengoba ucwaningo luye lwaqhubeka, izici zobuningi zomdlavuza oyinhloko we-colonectal, okuhlanganisa uhlobo lomdlavuza olufana neseli, i-immune, i-stromal, noma i-microbial, kuye kwabikwa ukuthi zihlobene kakhulu nemiphumela yezokwelapha, kodwa kusenokuqonda okulinganiselwe kokuthi ukusebenzisana kwazo kuthinta kanjani imiphumela yesiguli.
Ukuze kuhlolwe ubudlelwano phakathi kobunzima be-phenotypic kanye nomphumela, ithimba labacwaningi abavela eSidra Institute of Medical Research eQatar lisanda kuthuthukisa futhi laqinisekisa amaphuzu ahlanganisiwe (mICroScore) akhomba iqembu leziguli ezinezinga elihle lokusinda ngokuhlanganisa izici ze-microbiome kanye nama-immune rejection constants (ICR). Ithimba lenze ukuhlaziywa okuphelele kwe-genomic kwamasampula amasha aqandisiwe avela ezigulini ezingu-348 ezinomdlavuza we-colorectal oyinhloko, okuhlanganisa ukulandelana kwe-RNA kwamathumba kanye nezicubu ze-colorectal ezinempilo ezihambisanayo, ukulandelana kwe-exome ephelele, i-deep T-cell receptor kanye ne-16S bacterial rRNA gene sequencing, okunezelwe ukulandelana kwe-genome ye-tumor ephelele ukuze kuchazwe kabanzi i-microbiome. Ucwaningo lushicilelwe ku-Nature Medicine njenge-“An integrated tumor, immune and microbiome atlas of colon cancer”.

Isihloko esishicilelwe ku-Nature Medicine
Ukubuka konke kwe-AC-ICAM
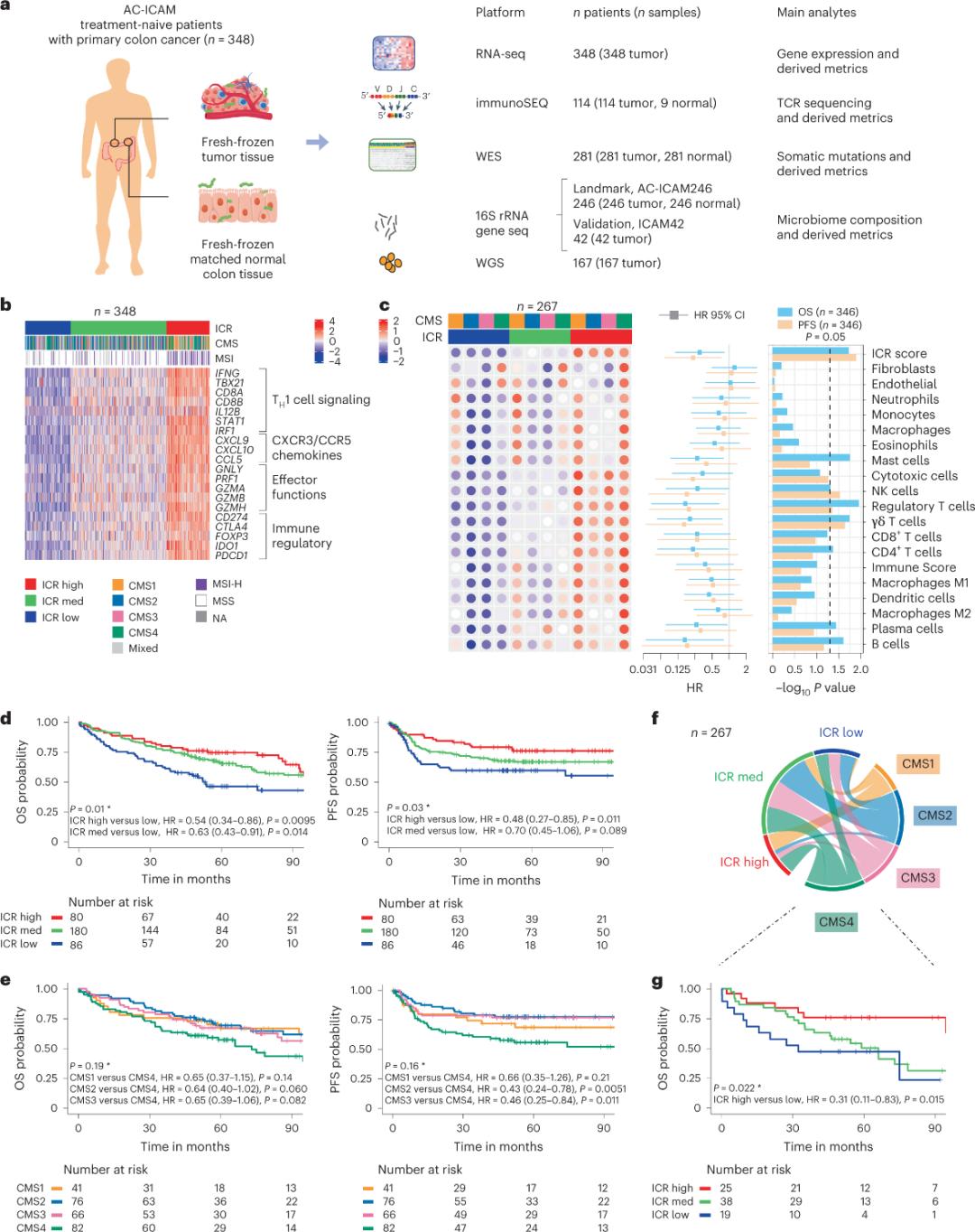
Abacwaningi basebenzise ipulatifomu ye-genomic ye-orthogonal ukuhlaziya amasampula esimila esiqandisiwe esisha futhi bafanisa izicubu zamathumbu aphilile eduze (ama-tumor-normal pairs) ezivela ezigulini ezine-histologic diagnosis yomdlavuza wamathumbu ngaphandle kokwelashwa okuhlelekile. Ngokusekelwe ekulandeleni kwe-whole-exome (WES), ukulawulwa kwekhwalithi yedatha ye-RNA-seq, kanye nokuhlolwa kwezindlela zokufakwa, idatha ye-genomic evela ezigulini ezingu-348 yagcinwa futhi yasetshenziselwa ukuhlaziywa okulandelayo ngokulandelela okuphakathi kweminyaka engu-4.6. Ithimba locwaningo laqamba lo mthombo ngokuthi i-Sidra-LUMC AC-ICAM: Imephu kanye nomhlahlandlela wokuxhumana kwe-immune-cancer-microbiome (Isithombe 1).
Ukuhlukaniswa kwama-molecule kusetshenziswa i-ICR
Bethatha isethi yezimpawu zofuzo zomzimba ezisetshenziswayo ukuze kuhlolwe umdlavuza okuqhubekayo, okubizwa ngokuthi i-immunoconstant of rejection (ICR), ithimba locwaningo lenze ngcono i-ICR ngokuyifinyeza ibe yiphaneli yezakhi zofuzo ezingama-20 ehlanganisa izinhlobo ezahlukene zomdlavuza, okuhlanganisa i-melanoma, umdlavuza wesinye, kanye nomdlavuza webele. I-ICR iphinde yahlotshaniswa nempendulo ye-immunotherapy ezinhlotsheni ezahlukene zomdlavuza, okuhlanganisa nomdlavuza webele.
Okokuqala, abacwaningi baqinisekisile isiginesha se-ICR seqembu le-AC-ICAM, besebenzisa indlela yokuhlukanisa i-ICR gene ukuze bahlukanise iqembu libe amaqoqo amathathu/ama-subtypes amasosha omzimba: i-ICR ephezulu (ama-hot tumors), i-ICR ephakathi nendawo kanye ne-ICR ephansi (ama-cold tumors) (Isithombe 1b). Abacwaningi bachaze ukuthambekela kokuzivikela komzimba okuhlotshaniswa nama-subtypes amangqamuzana avumelanayo (i-CMS), ukuhlukaniswa okusekelwe ku-transcriptome komdlavuza wamathumbu. Izigaba ze-CMS zazihlanganisa i-CMS1/immune, i-CMS2/canonical, i-CMS3/metabolic kanye ne-CMS4/mesenchymal. Ukuhlaziywa kubonise ukuthi amaphuzu e-ICR ayehlobene kabi nezindlela ezithile zamaseli omdlavuza kuzo zonke izinhlobo ze-CMS, futhi ukuhlangana okuhle nezindlela ezihlobene ne-immunosuppressive kanye ne-stromal kwabonwa kuphela kuma-tumors e-CMS4.
Kuwo wonke ama-CMS, ubuningi bamaseli e-natural killer (NK) kanye namaseli e-T babuphezulu kakhulu kuma-subtypes e-ICR aphezulu omzimba, kanye nokwehluka okukhulu kwamanye ama-subset e-leukocyte (Isithombe 1c). Ama-subtypes e-ICR immune ayene-OS ne-PFS ehlukene, kanye nokwanda okuqhubekayo kwe-ICR kusuka phansi kuya phezulu (Isithombe 1d), okuqinisekisa indima yokubikezela ye-ICR kumdlavuza we-colorectal.
Umfanekiso 1. Umklamo wocwaningo lwe-AC-ICAM, isiginesha yezakhi zofuzo ezihlobene nokuzivikela komzimba, izinhlobo ezincane zokuzivikela komzimba kanye nama-molecule kanye nokusinda.
I-ICR ibamba amaseli e-T akhuliswe ngamathumba futhi akhuliswe nge-clonal
Kubikwe ukuthi yingcosana yamaseli e-T angena ezicutshini zesimila akhethekile kuma-antigen esimila (ngaphansi kuka-10%). Ngakho-ke, iningi lamaseli e-T angaphakathi kwesimila abizwa ngokuthi amaseli e-T aqaphile (amaseli e-T aqaphile). Ubudlelwano obuqine kakhulu nenani lamaseli e-T avamile anama-TCR akhiqizayo bubonwe kumaseli e-stromal kanye nama-leukocyte subpopulations (atholwe yi-RNA-seq), angasetshenziswa ukulinganisa amaseli e-T subpopulations (Isithombe 2a). Kumaqoqo e-ICR (ngokuphelele kanye nokuhlukaniswa kwe-CMS), i-clonality ephezulu kakhulu yama-immune SEQ TCR abonwe kumaqembu e-ICR-high kanye ne-CMS subtype CMS1/immune (Isithombe 2c), enengxenye ephezulu kakhulu yama-tumors aphezulu e-ICR. Kusetshenziswa i-transcriptome yonke (amajini angu-18,270), amajini ayisithupha e-ICR (IFNG, STAT1, IRF1, CCL5, GZMA, kanye ne-CXCL10) ayephakathi kwamajini ayishumi aphezulu ahlotshaniswa kahle ne-TCR immune SEQ clonality (Isithombe 2d). Ukwakheka kwe-ImmunoSEQ TCR kuhlobene kakhulu nezakhi zofuzo eziningi ze-ICR kunokuhlangana okubonwe kusetshenziswa izimpawu ze-CD8+ ezisabela ku-tumor (Isithombe 2f no-2g). Ekuphetheni, ukuhlaziywa okungenhla kusikisela ukuthi isiginesha se-ICR sibamba ukuba khona kwamaseli e-T acebile nge-tumor, akhuliswe nge-clonal futhi singachaza imiphumela yaso yokubikezela.
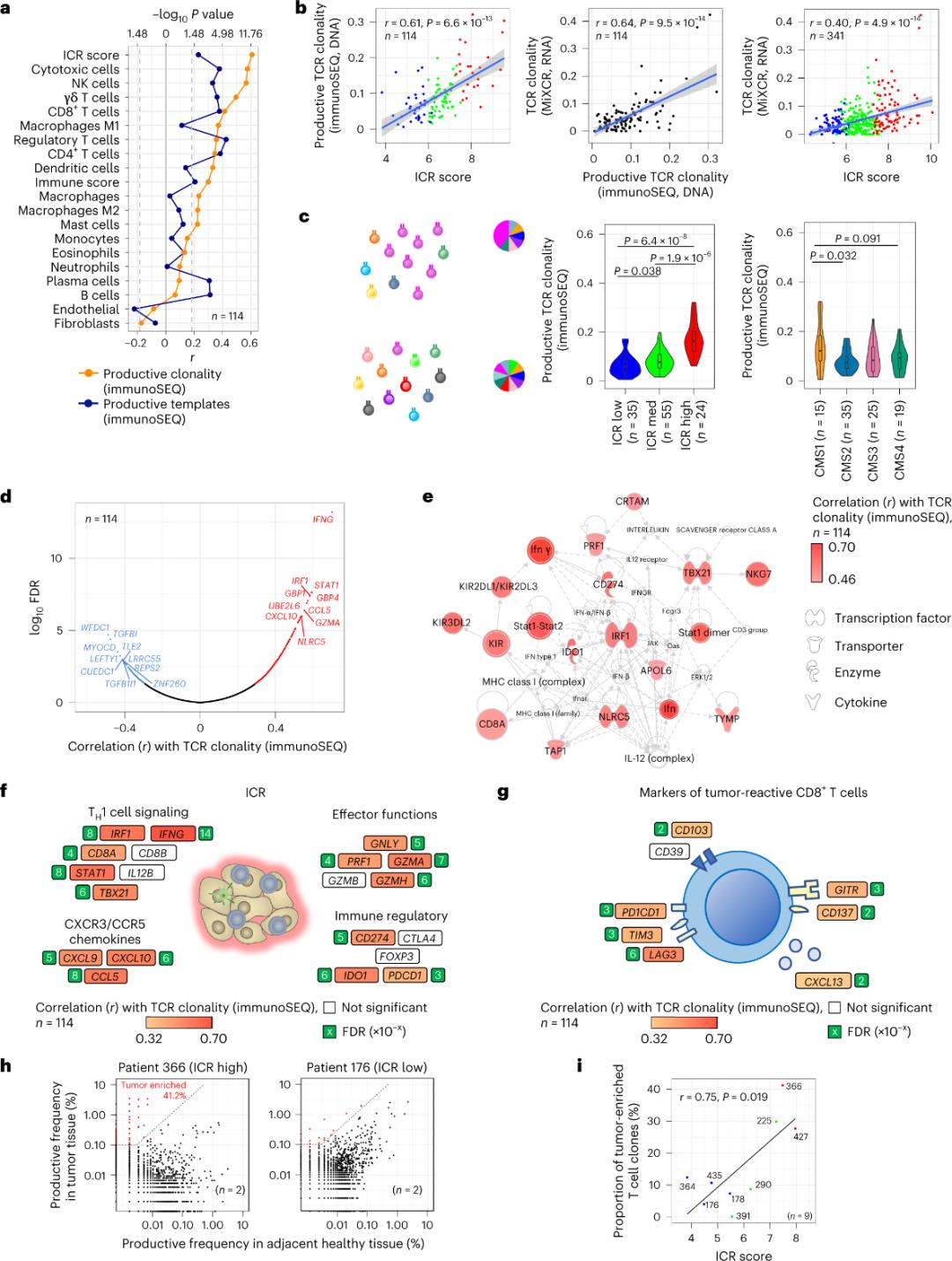
Umfanekiso 2. Izilinganiso ze-TCR kanye nokuhlobana nezakhi zofuzo ezihlobene nokuzivikela komzimba, izinhlobo ezincane zokuzivikela komzimba kanye nama-molecule.
Ukwakheka kwama-microbiome ezicutshini zomdlavuza wamathumbu aphilile kanye nomdlavuza wamathumbu
Abacwaningi benze ukulandelelana kwe-16S rRNA besebenzisa i-DNA ekhishwe esimila esifanelanayo kanye nezicubu zamathumbu aphilile ezigulini ezingu-246 (Isithombe 3a). Ukuze kuqinisekiswe, abacwaningi baphinde bahlaziya idatha yokulandelelana kwezakhi zofuzo ze-16S rRNA ezivela kumasampula engeziwe angu-42 esimila angazange afane ne-DNA evamile etholakalayo ukuze ihlaziywe. Okokuqala, abacwaningi baqhathanisa ubuningi bezitshalo phakathi kwezimila ezifanelanayo kanye nezicubu zamathumbu aphilile. I-Clostridium perfringens yanda kakhulu ezimila uma kuqhathaniswa namasampula aphilile (Isithombe 3a-3d). Kwakungekho mehluko obalulekile ekuhlukeni kwe-alpha (ukuhlukahluka kanye nobuningi bezinhlobo kusampula eyodwa) phakathi kwamasampula esimila kanye namasampula aphilile, futhi ukwehla okuncane kokuhlukahluka kwamagciwane kwabonwa ezimila ezisezingeni eliphezulu le-ICR uma kuqhathaniswa nezimila eziphansi ze-ICR.
Ukuze bathole ubudlelwano obufanele emitholampilo phakathi kwamaphrofayili e-microbial kanye nemiphumela yezokwelapha, abacwaningi bahlose ukusebenzisa idatha yokulandelana kwezakhi zofuzo ze-16S rRNA ukuze bathole izici ze-microbiome ezibikezela ukusinda. Ku-AC-ICAM246, abacwaningi basebenzise imodeli yokuguqulwa kwe-OS Cox ekhethe izici ezingu-41 ezinama-coefficients angewona awero (ahlotshaniswa nengozi yokufa ehlukene), abizwa ngokuthi ama-MBR classifiers (Isithombe 3f).
Kuleli qembu lokuqeqesha (ICAM246), amaphuzu aphansi e-MBR (MBR<0, i-MBR ephansi) ahlotshaniswa nengozi ephansi kakhulu yokufa (85%). Abacwaningi baqinisekisile ubudlelwano phakathi kwe-MBR ephansi (ingozi) kanye ne-OS ende kuma-cohorts amabili aqinisekiswe ngokuzimela (ICAM42 kanye ne-TCGA-COAD). (Isithombe 3) Ucwaningo lubonise ubudlelwano obuqinile phakathi kwe-endogastric cocci kanye namaphuzu e-MBR, afanayo esimila kanye nezicubu zamathumbu aphilile.
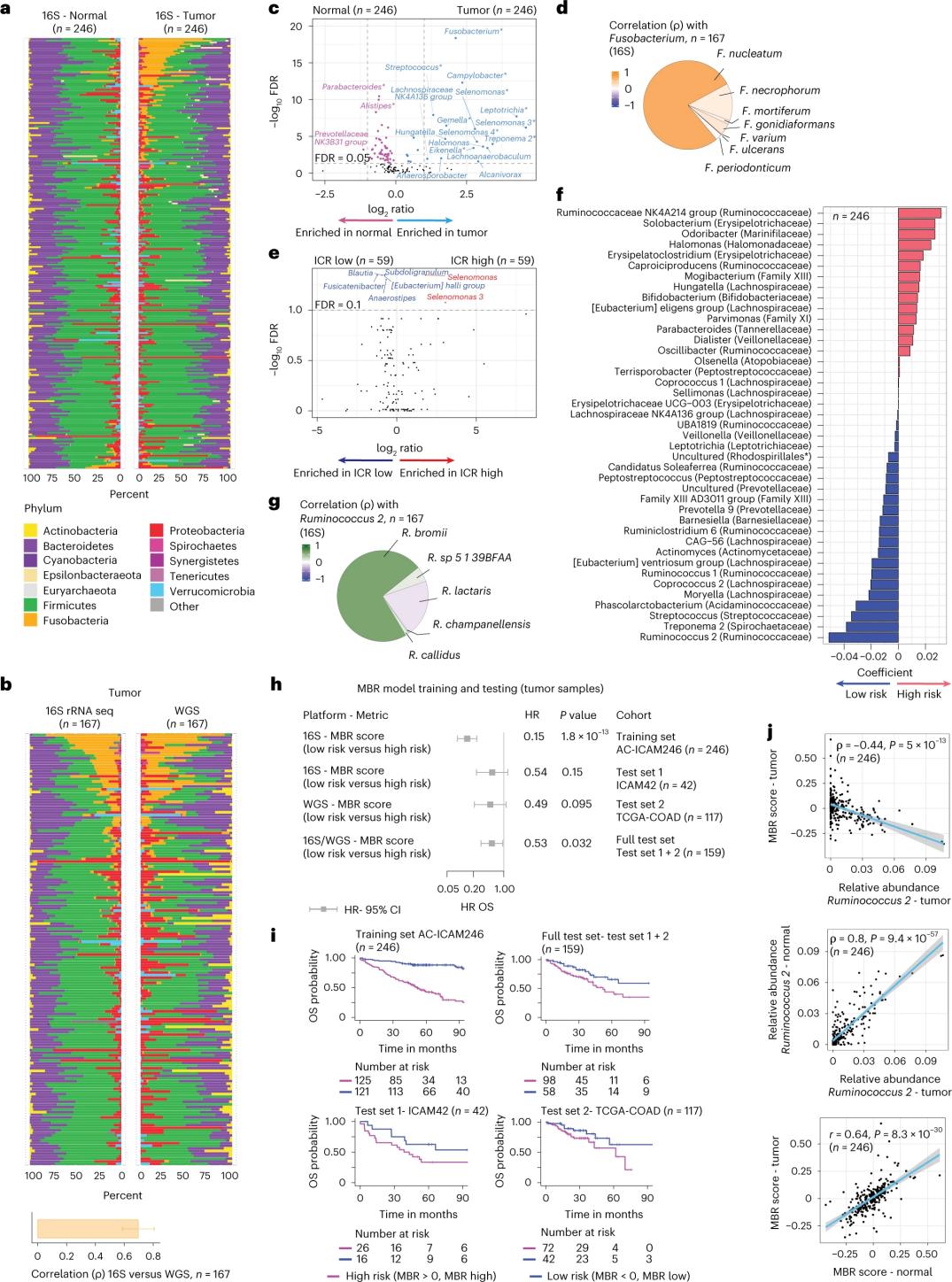
Isithombe 3. I-microbiome esimila nasezicutshini ezinempilo kanye nobudlelwano ne-ICR kanye nokusinda kwesiguli.
Isiphetho
Indlela ye-multi-omics esetshenziswe kulolu cwaningo ivumela ukutholwa okuphelele kanye nokuhlaziywa kwesiginesha yama-molecule yempendulo yomzimba kumdlavuza we-colorectal futhi yembula ukusebenzisana phakathi kwe-microbiome kanye nesistimu yomzimba. Ukulandelana okujulile kwe-TCR kwezicubu ezimilayo kanye nezicubu ezinempilo kwembule ukuthi umphumela wokubikezela we-ICR ungase ube ngenxa yekhono layo lokubamba ama-T cell clones acebile ngesimila futhi mhlawumbe ane-antigen ethile yesimila.
Ngokuhlaziya ukwakheka kwe-tumor microbiome kusetshenziswa ukulandelana kwezakhi zofuzo ze-16S rRNA kumasampula e-AC-ICAM, ithimba lithole isiginesha ye-microbiome (i-MBR risk score) enenani eliqinile lokubikezela. Nakuba lesi siginesha sithathwe kumasampula e-tumor, kwakukhona ukuhlobana okuqinile phakathi kwe-colorectum enempilo kanye ne-tumor MBR risk score, okuphakamisa ukuthi lesi siginesha singabamba ukwakheka kwe-gut microbiome yeziguli. Ngokuhlanganisa amaphuzu e-ICR kanye ne-MBR, kwakungenzeka ukuhlonza nokuqinisekisa i-biomarker yabafundi be-multi-omic ebikezela ukusinda ezigulini ezinomdlavuza we-colon. Isethi yedatha ye-multi-omic yocwaningo inikeza insiza yokuqonda kangcono i-biology yomdlavuza we-colon nokusiza ukuthola izindlela zokwelapha ezenzelwe wena.
Isikhathi sokuthunyelwe: Juni-15-2023
 中文网站
中文网站